Contact
hokumura_at_ims.ac.jp
Please replace the “_at_” with @
Introduction of Research
Biomolecules such as proteins and peptides have complicated free-energy landscape with many local minima. The conventional canonical-ensemble molecular dynamics (MD) simulations tend to get trapped in a few of the local-minimum states. To overcome these difficulties, we have proposed new generalized-ensemble algorithms, such as replica-permutation method. We apply these methods to reveal folding processes of some proteins and peptides.
We are also interested in neurodegenerative diseases that are caused by protein aggregates such as oligomers and amyloid fibrils. To understand formation of these protein aggregates, we perform replica-permutation MD simulations of these systems.
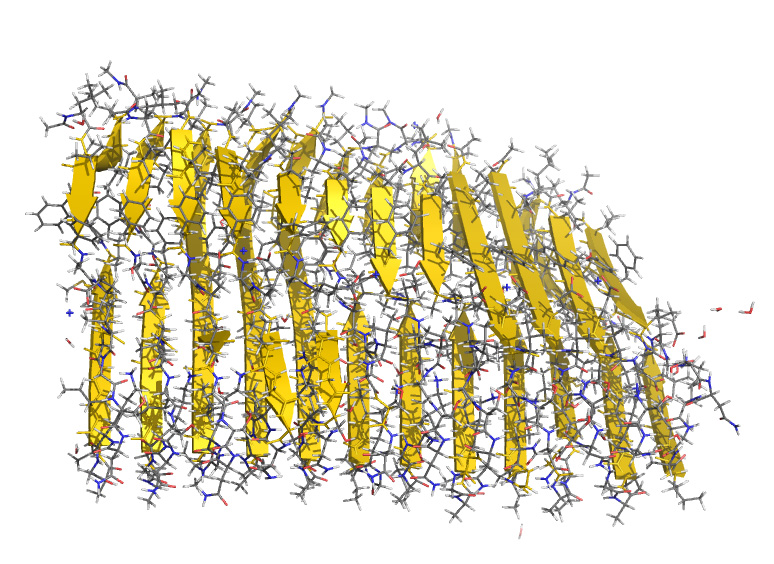 Amyloid fibril of amyloid-β peptides.
Amyloid fibril of amyloid-β peptides.