Remdesivir and favipiravir are currently attracting attention as therapeutic agents against COVID-19. These drugs are thought to act on the RNA-dependent RNA polymerase, a protein that replicates RNA by taking up nucleotides such as adenosine triphosphate (ATP). Remdesivir and favipiravir are expected to be taken up by the RNA polymerase instead of the nucleotides and inhibit RNA replication. However, it was not known how remdesivir, favipiravir, and ATP are taken up by the RNA polymerase.
The research group performed three types of molecular dynamics simulations in which 100 molecules of either remdesivir, favipiravir, or ATP were placed around the RNA polymerase. These molecules have phosphate groups, and it was found that the negative charges of the phosphate groups are attracted to the positive charges of the magnesium ions at the binding site of the RNA polymerase. The research group also identified three main paths in which these drugs reach the binding site. The RNA polymerase has two regions in which the lysine residues are arranged in a line. They found that these lysine residues correspond to two of the three paths. They succeeded in observing that the lysine residues attract phosphate groups of these drugs and ATP and sequentially pass them to the next lysine residue. In this way, it is revealed that the lysine residues transport the drugs and nucleotides to the binding site like a bucket brigade. In the third path, the drugs and nucleotides reach the binding site directly without contact with any amino acid residue. These results indicate that the RNA polymerase of SARS-CoV-2 utilizes electrostatic interaction to take up the nucleotides such as ATP and the drugs such as remdesivir and favipiravir. It is also found that to increase the uptake efficiency, the RNA polymerase arranges lysine residues in a line toward the binding site as if spreading tentacles, and carries them to the binding site like a bucket brigade.
Furthermore, it was found that the probability of uptake into the RNA polymerase binding site tends to be higher for favipiravir than for ATP, and for remdesivir than for favipiravir, although within statistical errors. However, further calculations are needed to make a definite determination.
This research has clarified for the first time in the world the mechanism by which the RNA polymerase of SARS-CoV-2 efficiently takes up nucleotides such as ATP for RNA replication. It was also revealed that the drugs such as remdesivir and favipiravir are also taken up by RNA polymerase by the same mechanism. This finding is expected to contribute to the development of more effective drugs to suppress the replication of SARS-CoV-2 in the future. Furthermore, it is known that lysine residues are arranged in a line not only in SARS-CoV-2 but also in SARS-CoV. Therefore, the findings of this study may contribute to the development of drugs against other viruses that have similar RNA polymerases.
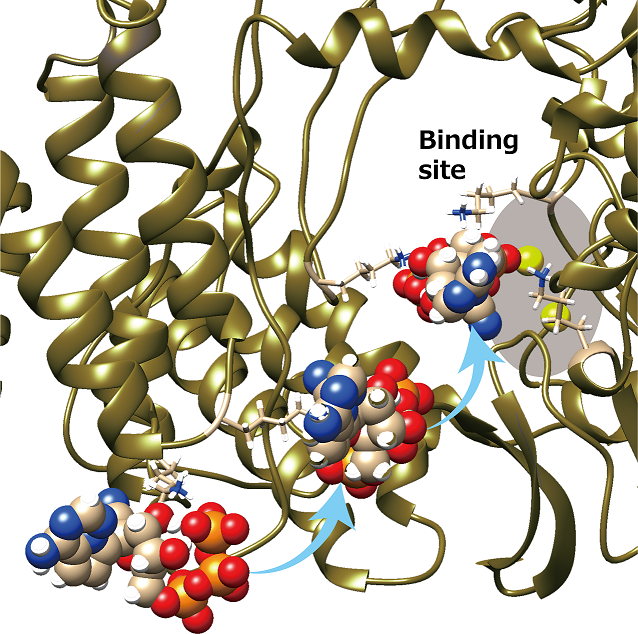
Path of remdesivir
Remdesivir (shown in the sphere model) is transferred to two magnesium ions (yellow-green spheres) at the binding site of the RNA polymerase (shown in the ribbon model) while being passed from one lysine residue (shown in the stick model) to another. We can see that the lysine residues transfer remdesivir as if it were a bucket brigade.
Here is a movie of a molecular dynamics simulation of remdesivir being transferred to the binding site of the RNA polymerase of SARS-CoV-2. The lysine residues transfer remdesivir like a bucket brigade.
Information of the paper
Authors: Shoichi Tanimoto, Satoru G. Itoh, and Hisashi Okumura
Journal Name: Biophysical Journal
Journal Title: ““Bucket brigade” using lysine residues in RNA-dependent RNA polymerase of SARS-CoV-2”
Research Group
Shoichi Tanimoto, Satoru G. Itoh, and Hisashi Okumura
Okumura Group, Institute for Molecular Science (IMS), Exploratory Research Center on Life and Living Systems (ExCELLS), National Institutes of Natural Sciences, Okazaki.
Contact:
Hisashi OKUMURA
Exploratory Research Center on Life and Living Systems (ExCELLS)
Institute for Molecular Science (IMS)
National Institutes of Natural Sciences, Okazaki.
TEL:0564-55-7277 FAX:0564-55-7025
E-mail:hokumura_at_ims.ac.jp
(Please replace the “_at_” with @)
Press contact
Exploratory Research Center on Life and Living Systems, National Institutes of Natural Sciences
TEL: +81-564-59-5201 FAX: +81-564-59-5202
E-mail: press_at_excells.orion.ac.jp
Public Relations, Research Enhancement Strategy Office, Institute for Molecular Science, National Institutes of Natural Sciences
TEL: +81-564-55-7209 FAX: +81-564-55-7374
E-mail: press_at_ims.ac.jp
(Please replace the “_at_” with @)

