Contact
yago_at_nips.ac.jp
Please replace the “_at_” with @
Introduction of Research
Spatiotemporal transcriptome regulations are essential for the proper construction of brain structure and function. Comprehensive analyses of the dynamics and the architecture of transcriptome in both wild and diseased animal models also lead to understanding the molecular causality of the human neuropsychiatric disease. Currently, our group examines the spatiotemporal transcriptome dynamics using the primate brain to identify the spatiotemporal-specific modulating genes from macro-scale to single-cell levels. Through this study, we aim to identify the molecular dynamics and trajectories between proper and atypical brain gene expressional networks. Additionally, we perform a massive population genetic analysis for the neuropsychiatric related genes in primates to identify an individual that has a spontaneous loss-of-functional (LoF) mutation in the neuropsychiatric genes and aim to make primate disease models for the neuropsychiatric study.
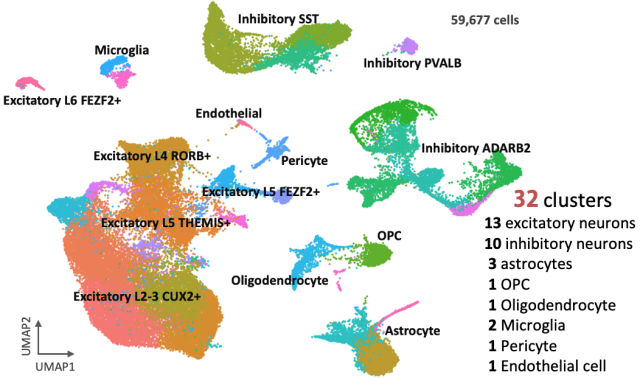 Identification of various types of brain cells in the marmoset prefrontal cortex via single-nucleus RNA-sequence experiment
Identification of various types of brain cells in the marmoset prefrontal cortex via single-nucleus RNA-sequence experiment