Researchers at the Exploratory Research Center on Life and Living Systems (ExCELLS), Nagoya University, and the Institute for Glyco-core Research (iGCORE) of Gifu University made significant progress in understanding the binding mechanism of a member of cell-cell adhesion molecules called CELSR cadherins in solution. Utilizing single-molecule fluorescence microscopy and high-speed atomic force microscopy (HS-AFM), these researchers found that two CELSR cadherin molecules extending from apposed cells interact with a twisted conformation to link the neighboring cells. These finding provide crucial insights into the intricate mechanism by which CELSR cadherins mediate adhesion between cells. Elucidating the detailed molecular mechanism by which CELSR cadherins recognize each other will help in comprehending development of the sophisticated human body plan, and offer insights into diseases caused by defects in CELSR cadherin function.
Our bodies comprise different tissues and organs, which are composed of many cells that must adhere to form functional higher order structures. This adherence is facilitated by specialized proteins called cell-cell adhesion molecules, which extend from neighboring cells and link to each other. One class of cell-cell adhesion molecules is the Cadherin EGF LAG seven-pass G-type receptor (CELSR) cadherins. These proteins play important roles in forming tissues that have a specific shape and pattern of cells. To date, we do not fully understand CELSR cadherin structure and function
Shigetaka Nishiguchi of ExCELLS, Rinshi S. Kasai of iGCORE at Gifu University, and Takayuki Uchihashi of ExCELLS and Nagoya University utilized single-molecule fluorescence microscopy and high-speed atomic force microscopy (HS-AFM) to elucidate the mechanism of CELSR cadherin dimerization. Researchers constructed two distinct fluorescently tagged CELSR cadherins (CELSR2-mGFP and CELSR2-Halo) and observed the interfaces between neighboring cells expressing CELSR2-mGFP or CELSR2-Halo using single-molecule fluorescence microscopy (Fig. A). Their results demonstrated that colocalization fluorescence spots of two differently tagged CELSR cadherins at cell-cell interfaces were sustained above the long distances (~1 μm) and time (~1 sec), indicating that CELSR cadherins form specific and stable dimers to connect neighboring cells to form a minimal adhesion unit. Next, they used HS-AFM to obtain high-resolution images of purified CELSR cadherins in solution at the nanometer scale (Fig. B). The images revealed that the extracellular domains (ectodomains) of CELSR cadherin that connect cells exhibited strand- and globule-like portions composed of cadherin ectodomains and non-cadherin ectodomains, respectively. HS-AFM also revealed that the two CELSR cadherins bind through strand-like structures in an antiparallel orientation, consistent with the binding complex of CELSR cadherins between cells observed using single-molecule fluorescence microscopy. These researchers applied localization analysis of HS-AFM images previously developed by other researchers to improve the spatial resolution of their HS-AFM images and obtain higher-resolution images of the structure. Their analysis showed that the strand-like portions of the eight cadherin ectodomains of the CELSR cadherins overlapped entirely in a twisted manner. These researchers have also demonstrated that the fourth cadherin ectodomain from the membrane-distal side of CELSR cadherin is especially important for CELSR cadherin interactions using a bead aggregation assay, which can evaluate CELSR cadherin binding activity.
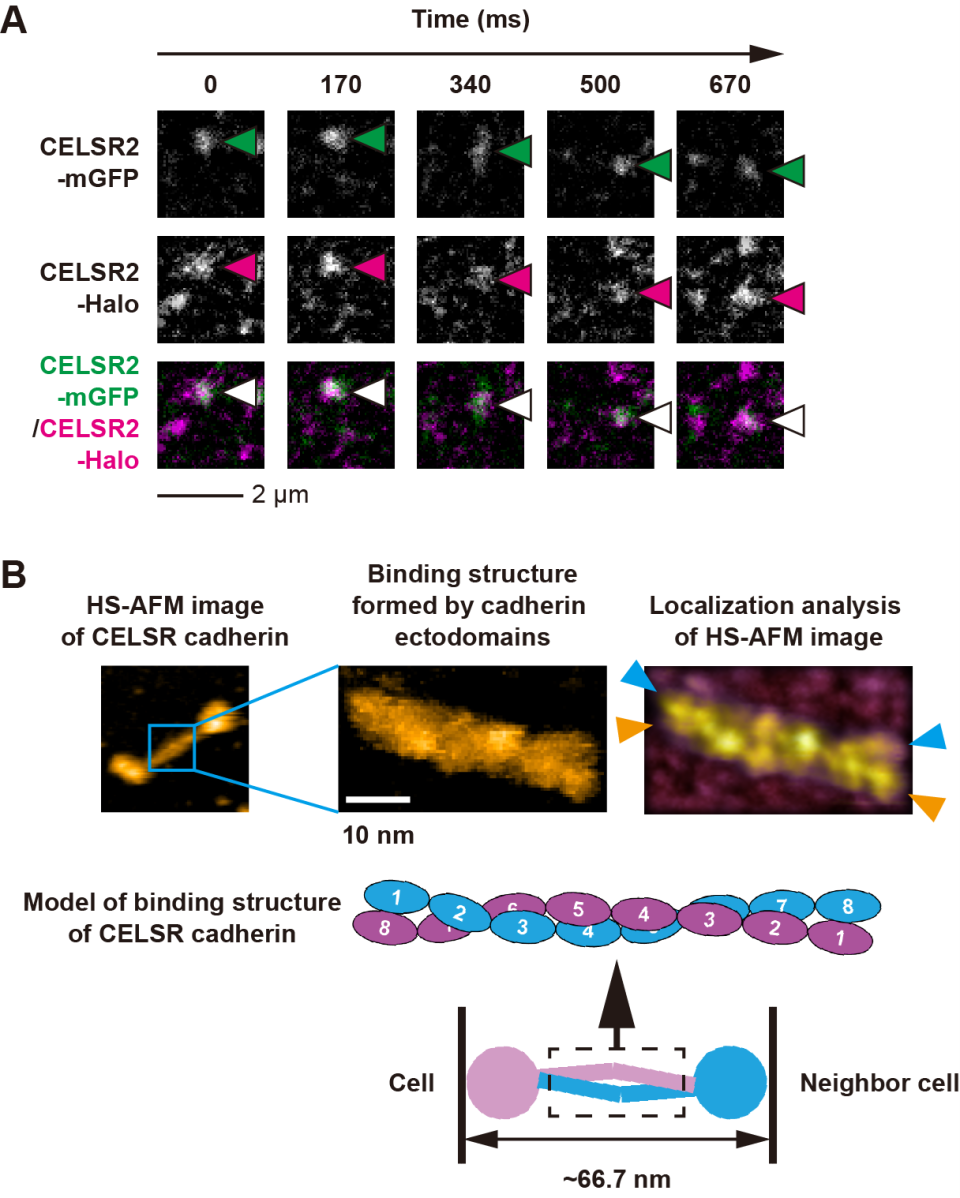 Figure 1. Binding mechanism of CELSR cadherin.
Figure 1. Binding mechanism of CELSR cadherin.
(A) Single-molecule fluorescence microscopy images of CELSR cadherins from adjacent cells. Two differently tagged CELSR cadherins (CELSR2-mGFP and CELSR2-Halo; indicated with arrows) colocalize between cells. (B) High-speed atomic force microscopy (HS-AFM) images of a complex of two interacting CELSR cadherins. A twisted conformation in the strand-like portions of the cadherin ectodomains is visualized using localization analysis of the HS-AFM images. Blue and orange colored arrowheads indicate the tip of each CELSR cadherin.
Interestingly, the overlapping of eight cadherin ectodomains in the binding complex reported in this study is the most extensive interaction described to date relative to other cadherin members, and the CELSR cadherin complex is larger (~66.7 nm) than the typical extracellular space distance in the cell-cell adhesion area of epithelial cells (15–25 nm). Researchers have hypothesized that the large binding complex formed by CELSR cadherins may act as a physical spacer between cells, allowing space for small extracellular vesicles containing messenger molecules, and regulating polarity-dependent tissue formation, which directly impacts the patterning of cells. These findings provide new insights into the complex mechanisms by which CELSR cadherins mediate cellular adhesion. Although further detailed analyses are necessary to fully understand the physiological role of the binding structure of the CELSR cadherins reported in this study, the results have important implications for the development of new therapies for diseases involving impaired cell-cell adhesion.
This work was supported by a Grant-in-Aid for Early-Career Scientists JP 22K14577 (to S.N.); a Grant-in-Aid for Scientific Research JP 17K07333, JP 18H05269, JP 18H05424, and JP 20H03225 (to R.S.K.); JP 21H00393, JP 21H01772, and JP 22K18943 (to T. U.) from the Japan Society for the Promotion of Science; and by the Joint Research of the Exploratory Research Center on Life and Living Systems.
Reference
Journal:
Proceedings of the National Academy of Sciences
Method of research:
Experimental study
Subject of research:
Cells
Airticle title:
Antiparallel dimer structure of CELSR cadherin in solution revealed by high-speed atomic force microscopy
Airticle publication date :
24-Apr-2023
Eurekalert! URL : https://www.eurekalert.org/news-releases/987377
Contact
Media contact :
Strategic Research Administration Office
Exploratory Research Center on Life and Living Systems (ExCELLS)
National Institutes of Natural Sciences
E-Mail: press_at_excells.orion.ac.jp
(Please replace the “_at_” with @)
Expert contacts :
NISHIGUCHI, Shigetaka
Graduate School of Engineering, Osaka University
( At the time of research: Exploratory Research Center on Life and Living Systems (ExCELLS) )
E-mail: nishiguchi.shigetaka.hyf_at_osaka-u.ac.jp
(Please replace the “_at_” with @)

